Tuberculosis diagnostics in the 21st century
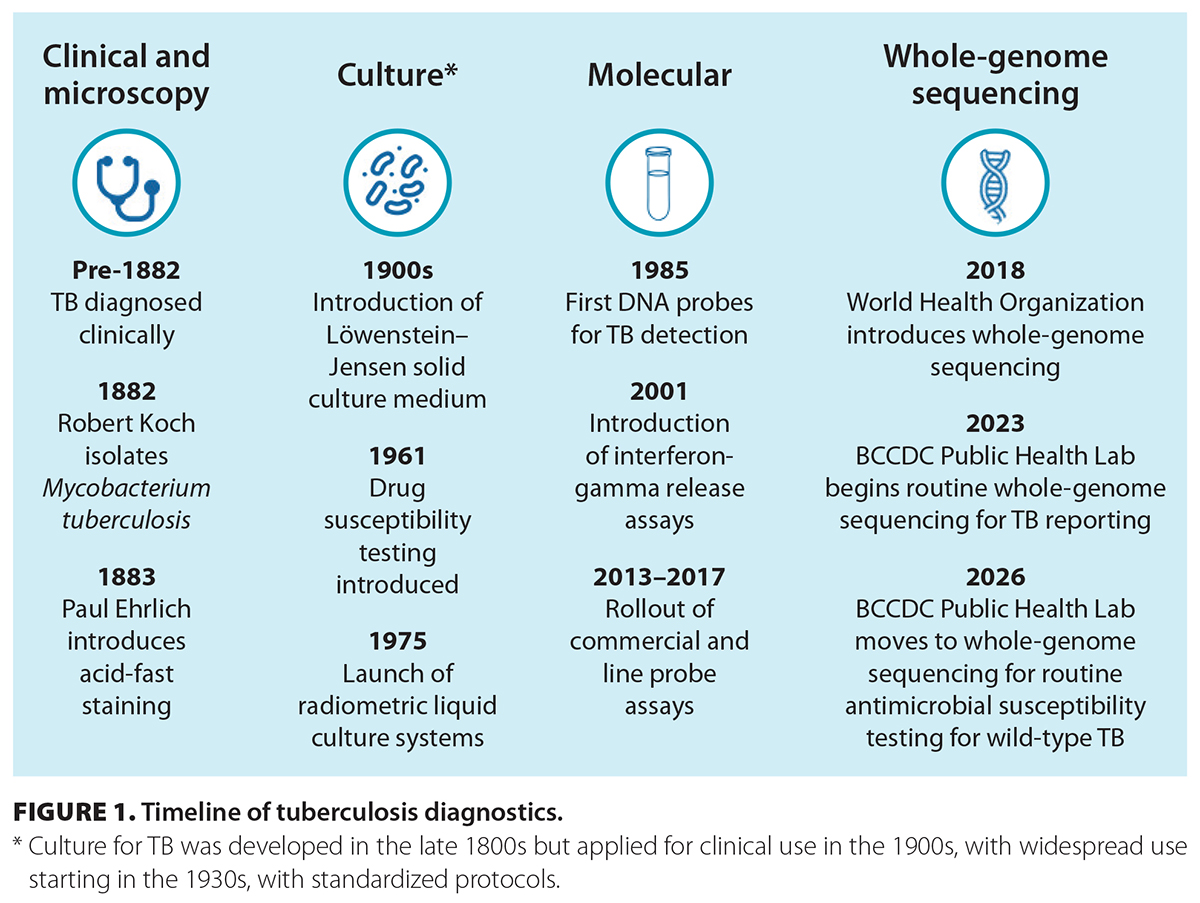 Tuberculosis (TB) diagnostics relied on clinical acumen until the late 19th century, when microscopy and culture were developed. This was followed by susceptibility testing, which was developed in parallel to antibiotic therapy. Recently, molecular techniques have been changing centuries-old methods [Figure 1]. Yet, TB remains the most common cause of infectious mortality, with close to 1.25 million deaths worldwide in 2024.[1] New laboratory techniques are key to reaching the elimination goal of less than 1 person per million with TB globally by 2050.[2]
Tuberculosis (TB) diagnostics relied on clinical acumen until the late 19th century, when microscopy and culture were developed. This was followed by susceptibility testing, which was developed in parallel to antibiotic therapy. Recently, molecular techniques have been changing centuries-old methods [Figure 1]. Yet, TB remains the most common cause of infectious mortality, with close to 1.25 million deaths worldwide in 2024.[1] New laboratory techniques are key to reaching the elimination goal of less than 1 person per million with TB globally by 2050.[2]
Molecular techniques provide faster and more accurate information about TB, including direct detection and sequencing for epidemiology and susceptibility prediction. This year marks a turning point in TB diagnostics. As of 5 January 2026, most susceptibility testing in British Columbia will rely on genomic prediction rather than phenotypic. This change speeds up the availability of results from 21 days to as few as 7 days following culture positivity.
Highlights of recent TB diagnostic developments
Direct detection by polymerase chain reaction
Polymerase chain reaction (PCR) was introduced in the 1980s and has been used widely for TB diagnostics since the early 2000s. This marked the first time it was possible to confirm TB without relying on culture. Early PCR had variable sensitivity, often lower than that of standard microscopy. In the early 2010s, improved sensitivity allowed for reliable and reproducible detection of TB from sputum. New GeneXpert testing reduces complexity and speeds turnaround time, enabling clinical teams to quickly be informed of positive specimens. The most notable advance in TB PCR diagnostics in the last 10 years is resistance detection for the antibiotics isoniazid and rifampin. These tests can help identify patients with multidrug-resistant TB to rapidly optimize treatment and reduce the risk of onward transmission.
Next-generation sequencing to identify nontuberculous mycobacteria species
While TB is the primary target of detection and treatment programs, nontuberculous mycobacteria (NTM) continue to cause disease and confuse TB diagnosis. Historic identification and differentiation of these organisms have relied on phenotypic and biochemical behavior, taking weeks to months. Molecular advances, particularly next-generation sequencing, enable the use of sequenced regions to classify NTM within 2 to 3 days of growth, often to the level of subspecies or variants. This technique can also identify TB and mixed infections, thereby avoiding the need for complex procedures to separate mixed bacterial populations. The assay used at the BCCDC Public Health Laboratory also identifies common resistance mutations. Rapid and accurate identification of NTM species allows TB to be ruled out and enables decisions about the clinical significance of the organism for the patient.
Whole-genome sequencing
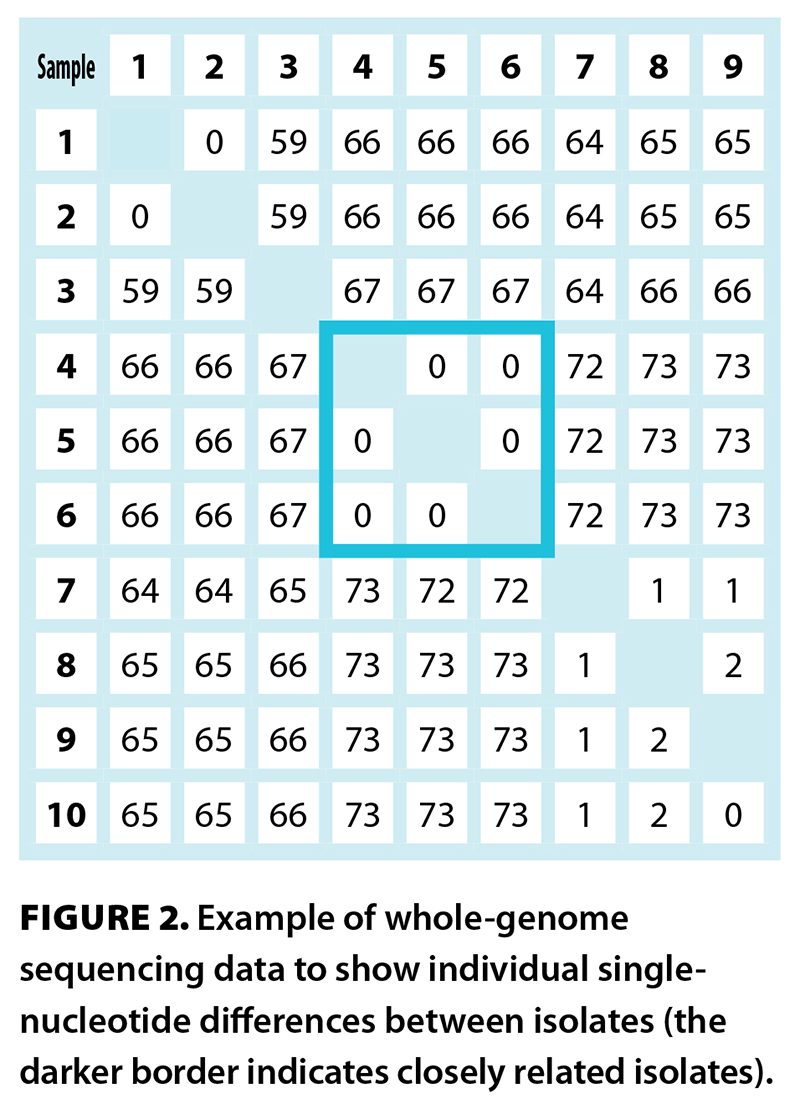 Since entering clinical use in 2009, whole-genome sequencing (WGS) has transformed medicine. Use for TB diagnostics began in 2012 to support cluster investigations, replacing older methods that relied on repeats and other short genetic sequences and lacked granularity. WGS allows the identification of single-nucleotide differences between strains, which can identify transmission events and uncover previously undetected transmission networks [Figure 2]. WGS was first used in a research capacity in BC in 2018 and was transitioned to regular clinical application in 2023—the BCCDC Public Health Laboratory was the first provincial laboratory to rely on WGS results. We now have over 2000 isolates sequenced, providing a rich data set to understand epidemiology and confirm accuracy. Techniques and workflows are continually refined to minimize time to definitive results and support traditional epidemiology by confirming cases and clusters for further investigation.
Since entering clinical use in 2009, whole-genome sequencing (WGS) has transformed medicine. Use for TB diagnostics began in 2012 to support cluster investigations, replacing older methods that relied on repeats and other short genetic sequences and lacked granularity. WGS allows the identification of single-nucleotide differences between strains, which can identify transmission events and uncover previously undetected transmission networks [Figure 2]. WGS was first used in a research capacity in BC in 2018 and was transitioned to regular clinical application in 2023—the BCCDC Public Health Laboratory was the first provincial laboratory to rely on WGS results. We now have over 2000 isolates sequenced, providing a rich data set to understand epidemiology and confirm accuracy. Techniques and workflows are continually refined to minimize time to definitive results and support traditional epidemiology by confirming cases and clusters for further investigation.
In addition, WGS offers the ability to predict susceptibility. Since 2018, the World Health Organization has collated and published resistance mutations, permitting bioinformatics teams to link sequences to phenotypic behavior. The advantages of WGS are that results can be made available in several days to weeks, and multiple antibiotics can be assessed with a single test. The BCCDC Public Health Laboratory has validated the first-line agents compared with phenotypic methods, finding excellent concordance. Negative predictive values for wild-type TB to first-line medications are 99.5% for isoniazid, 99.6% for pyrazinamide, 100% for ethambutol, and 100% for rifampin. Fully susceptible organisms are now reported with confidence within 10 days of detection, an improvement of 10 to 14 days.
All these advances rely on a large team of individuals with advanced skills—laboratory personnel, bioinformaticians, informatics specialists, front-line clinicians, epidemiologists, and many others all play key roles. We are grateful for the wonderful team working together to eliminate TB.
—Jennifer M. Grant, MDCM, FRCPC
Associate Medical Director and Program Head, Mycobacteriology, BCCDC Public Health Laboratory
—Victoria J. Cook, MD, FRCPC
Medical Head, Provincial Tuberculosis Services, BCCDC
—James Johnston, MD, MPH, FRCPC
Evaluation Lead, Provincial Tuberculosis Services, BCCDC
hidden
This article is the opinion of the BC Centre for Disease Control and has not been peer reviewed by the BCMJ Editorial Board.
 |
| This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License. |
References
1. World Health Organization. Tuberculosis. 13 November 2025. Accessed 15 January 2026. www.who.int/news-room/fact-sheets/detail/tuberculosis.
2. Cook VJ, Matsuba M. Provincial Tuberculosis Committee: Tuberculosis quality care and elimination plan 2024-2029. 2024. Accessed 15 January 2026. www.bccdc.ca/resource-gallery/Documents/%20and%20Forms/Guidelines%20and%20Manuals/TB/TB_Quality_Care_Elimination_Plan.pdf.
3. Carroll LN, Au AP, Detwiler LT, et al. Figure 4. Dendrogram. In: Visualization and analytics tools for infectious disease epidemiology: A systematic review. J Biomed Inform 2014;51:287-298. Accessed 15 January 2026. https://doi.org/10.1016/j.jbi.2014.04.006.

